Professional technical scientists and abundant experience in immune checkpoint drug development make Creative Biolabs a perfect partner to help our clients’ research in the immunological field. Currently, we provide high-quality services for immune checkpoint targeted small molecule drug design, particularly structure-based immune checkpoint targeted drug design for our global customers.
Immunotherapy has revolutionized cancer therapy and resulted in a paradigm shift in the war on cancer. Immune checkpoints play important roles in mediating immune tolerance and protecting tissues from collateral damage. Drugs targeting immune checkpoints such as immune checkpoint inhibitors have demonstrated considerable promise in the treatment of cancers. Until now, scientists have identified hundreds of molecules targeting several pathways. These drugs are classified into six groups according to function and cell of predominant expression: lymphoid inhibitors (LAG-3, TIM3, TIGIT), lymphoid stimulators (OX40, GITR, 4-1BB, ICOS), non-lymphoid inhibitors (IDO1, TGFβ, CD47), non-lymphoid stimulators (CD40), NK stimulators (KIR2R and IL15) and inhibitors (NKG2A).

The research-based pharmaceutical industry has increasingly employed modern medicinal chemistry methods, including molecular modeling. This field has progressed together with advances in biomolecular spectroscopic techniques such as X-ray crystallography and nuclear magnetic resonance (NMR), which have enabled striking progress in molecular and structural biology. Based on our deep understanding of these advanced drug design strategies, Creative Biolabs has extensive experience in offering custom services for successful immune checkpoint drug discovery to meet our clients' demands precisely.
Structure-based drug design (SDBB) methods (using three-dimensional structural information gathered from biological targets) are a prominent component of modern medicinal chemistry. In most cases, the protein structure used in the structure-based design process has been determined by X-ray crystallography. Molecular docking, structure-based virtual screening (SBVS), and Ligand-Based Virtual Screening (LBVS) are among the most frequently used SDBB strategies due to their wide range of applications in the analysis of molecular recognition events. Along with the prediction of the binding mode, SBVS provides a ranking of the docked molecules for the sole criterion for selecting promising molecules, or it can be combined with other evaluation methods.
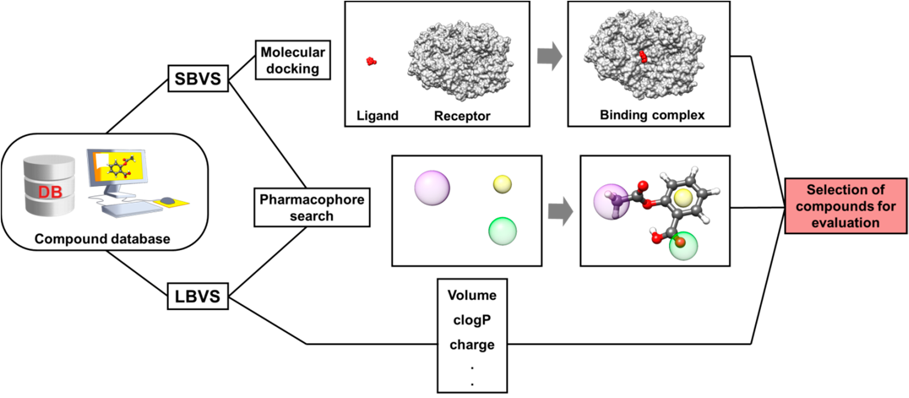 Fig.1. The SBVS and LBVS approach.1,2
Fig.1. The SBVS and LBVS approach.1,2
In general, X-ray crystallography is the preferred technique for elucidating the three-dimensional (3D) structure of target-small molecule ligand complexes. There are, however, several cases where NMR is indispensable. One obvious case is systems where crystallization fails due to the dynamic nature of the target. Other more classical cases involve targets that can be crystallized but where soaking or crystallization fails because the binding site is not accessible. Experienced scientists at Creative Biolabs are glad to apply our special expertise to offer NMR fragment-based immune checkpoint targeted small molecule drug discovery.
With years of experience in drug design and structural biology, Creative Biolabs is dedicated to offering structure-based immune checkpoint targeted drug design services to support the immunotherapy industry. We could use various technologies, such as X-ray crystallography, protein NMR spectroscopy, homology modeling, small molecule docking, to meet each requirement from our global customers. Please contact us for more detailed information to know how we can help.
References
All listed customized services & products are for research use only, not intended for pharmaceutical, diagnostic, therapeutic, or any in vivo human use.
USA
Tel:
Fax:
Email:
Copyright © 2026 Creative Biolabs. All Rights Reserved.